Stanford researchers make first direct observation of 3-D molecule folding in real time
All the crucial proteins in our bodies must fold into complex shapes to do their jobs. These snarled molecules grip other molecules to move them around, to speed up important chemical reactions or to grab onto our genes, turning them "on" and "off" to affect which proteins our cells make.
Recently, scientists have discovered that RNA can act a lot like a protein itself. RNA can form loopy bundles that shut genes down or start them up without the help of proteins. Since the discovery of these RNA clumps, called "riboswitches," in 2002, scientists have been striving to understand how they work and how they form. Now, researchers at Stanford University are looking closer than ever at how the three-dimensional twists and turns in a riboswitch come together by grabbing it and tugging it straight. By physically pulling on this loopy RNA, they have determined for the first time how a three-dimensional molecular structure folds, step by step.
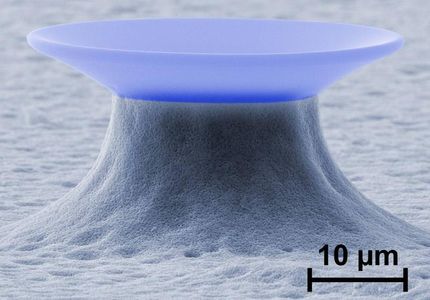
The researchers used a machine called an "optical trap" to grab and hold the ends of an RNA molecule with laser beams. Based on technology developed by Bell Labs researchers in 1986, the machine was designed by a team led by Steven Block, the Stanford W. Ascherman, M.D., Professor and a professor of applied physics and of biology. The optical trap allows them to hold the ends of the RNA tightly, so they can pull it pin-straight, then let it curl up again. In "Science", their paper, of which Block is senior author, describes the development of every loop and fold in one particular RNA riboswitch, and the energy it takes to form or straighten each one-an unprecedented achievement that opens the door for equally thorough studies of other molecules and their behaviors.
The researchers are the first to study the energy and folding behavior of a riboswitch in this detailed, physical way. More important, they are the first to use directly applied force to determine how a molecule makes a three-dimensional bundle, a tertiary structure. No other research has tracked the formation of such a complex structure, fold by fold.
Previous studies typically have used biochemical techniques rather than lasers, which can directly grab and tug the RNA. Biochemical techniques give less clear estimates of how molecules fold in real time. They often give a description of the molecule's average folding behavior, which must be interpreted by mathematical models. Crystallography-a technique involving freezing the molecule in place-provides a good picture of its shape, but not how it forms or the energy involved.
"What we're interested in is understanding, in a very fundamental way, how biomolecules take the shapes they do, and how they perform the functions they do," Block said. "No one has been able to explore in great detail tertiary structure yet." RNA riboswitches must have this tertiary structure to work.
"Most RNAs just make secondary [two-dimensional] structure. But the ones that really do stuff," he added, "those all have tertiary structure."
Weitere News aus dem Ressort Wissenschaft
Diese Produkte könnten Sie interessieren

DynaPro NanoStar II von Wyatt Technology
NanoStar II: DLS und SLS mit Touch-Bedienung
Größe, Partikelkonzentration und mehr für Proteine, Viren und andere Biomoleküle

AZURA Purifier + LH 2.1 von KNAUER
Präparative Flüssigkeitschromatografie - Neue Plattform für mehr Durchsatz
Damit sparen Sie Zeit und verbessern die Reproduzierbarkeit beim Aufreinigen

Holen Sie sich die Chemie-Branche in Ihren Posteingang
Mit dem Absenden des Formulars willigen Sie ein, dass Ihnen die LUMITOS AG den oder die oben ausgewählten Newsletter per E-Mail zusendet. Ihre Daten werden nicht an Dritte weitergegeben. Die Speicherung und Verarbeitung Ihrer Daten durch die LUMITOS AG erfolgt auf Basis unserer Datenschutzerklärung. LUMITOS darf Sie zum Zwecke der Werbung oder der Markt- und Meinungsforschung per E-Mail kontaktieren. Ihre Einwilligung können Sie jederzeit ohne Angabe von Gründen gegenüber der LUMITOS AG, Ernst-Augustin-Str. 2, 12489 Berlin oder per E-Mail unter widerruf@lumitos.com mit Wirkung für die Zukunft widerrufen. Zudem ist in jeder E-Mail ein Link zur Abbestellung des entsprechenden Newsletters enthalten.